Tutorials
Scalar Field Dataset: X-ray
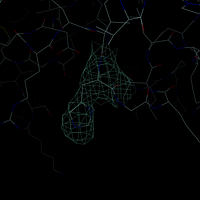
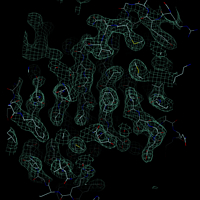
An electron density map used in x-ray crystallography is very easy to display with DINO. In contrast to other datasets, a few properties need to be defined during object creation (otherwise defaults will be used), such as center of the map (default {0,0,0}), the size (default 30) and the contour level (default 0). Appending the letter 's' to the contour level means sigma units.
Download this tutorial: scal_xray.tar.gz
// example using IIB subunit of the fructose transporter
// Schauder et. al. 1997
// preset CS so that script works
set CS .iib:15.CA
// load the structure dataset
load iib.pdb
// load the scalar field dataset
// (in this case a CCP4 electron density map)
load iib_2fofce.map -name m
// generate structure object and center on it
.iib new -name all
scene center [.iib.all]
// adjust the clipping planes
scene autoslab
// create a new map object, centered on structure object,
// contour level 1sigma unit, 20x20x20 grid units in size
.m new -name m -set center=[.iib.all],level=1.0s,size=20
.m.m set color=aquamarine
// to recenter the map on another point, select an atom
// with the mouse and then use
.m.m renew -set center=[$CS]
scene center [$CS]
// to restrict the map around certain regions, use the
// within operator
// create a new structure object, in this case
// only residue H15
.iib new -name h15 -sel rnum=15
// renew the map with a selection statement
// to cover only residue 15
.m.m renew -sel 2.2<>.iib.h15
// to remove the selection criteria use
.m.m renew -sel *
|